Tipping-Point Detection
Tipping-Point Detection (TPD) is a webserver for data analysis and visualization of tipping point detection. More about TPD
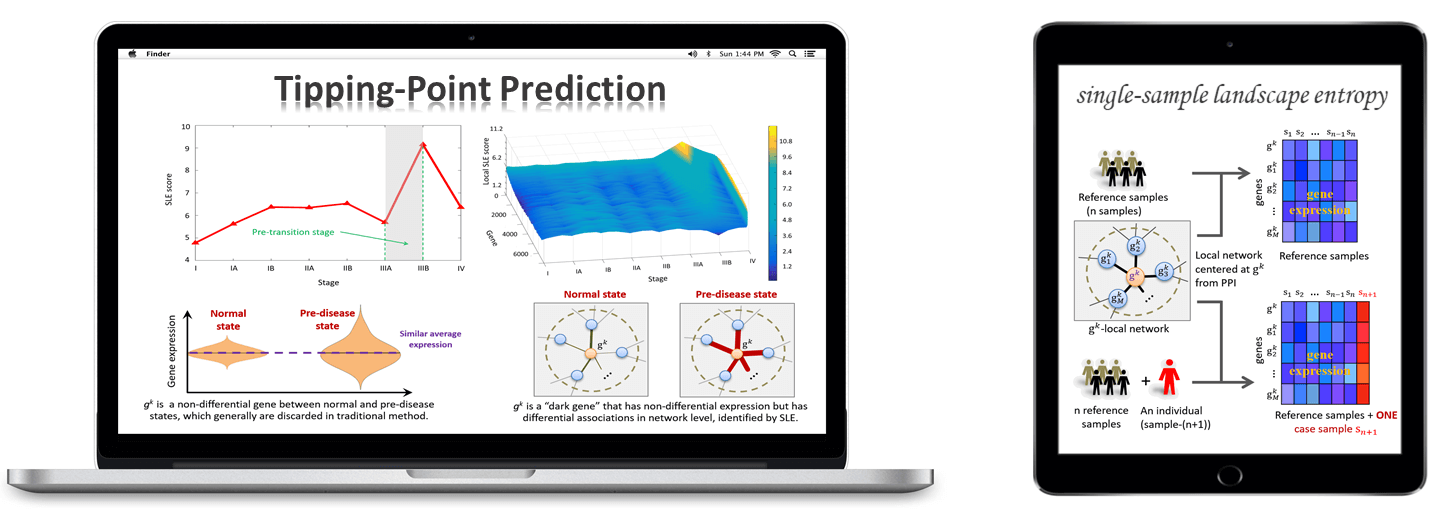
TPD: Tipping-Point Prediction.
The time evolution or dynamic change of many biological systems, such as disease progression and cell differentiation process, is not always smooth but occasionally abrupt, that is, there is a tipping-point or critical state during such a process at which the system state shifts irreversibly from one state (e.g.normal state) to another state (e.g. disease state), just before the critical transition. It is challenging and also important to predict such a critical state with the measured omics data, which is a key to achieve the predictive or preventive medicine. In this summary, we introduce a web service, the Tipping-Point Prediction (TPD), developed to effectively and rapidly identify the tipping point during the dynamical process of biological systems, and further its leading molecules or network. The web service is based on our computational method called sample-based local network entropy (SNE), which is a model-free data-driven approach with solid theoretical background, i.e., the dynamic network biomarker (DNB). Specifically, TPD is user-friendly and capable to explore the criticality of the dynamics from high-dimensional omics-data in terms of network entropy, thereby capturing not only the early-warning signals of the impending transition but also its leading network (DNB), which are intuitively a group of strong correlated and fluctuated molecules.
The details of the web service are as follows: The input of TPD should be the time-series or stage-course data (expression matrix) in CSV, XLS, TXT files. In the web tool, there is a flexible option for users to assign samples to certain time points. The web tool outputs multifarious visualized results including SNE scores and curves with suggested critical point, the identified key genes (or leading genes) and their information, DNB (the leading network that may drive the critical transition), the dynamical process of the leading network, the survival analysis based on SNE score that may help to identify dark genes (non-differential in terms of expression but differential in terms of network entropy, which may play important roles during the dynamic evolution). Thus, the web tool offers significantly enhanced display of the data and results.
Browser compatibility: TPD web tool is user-friendly for common browsers in operation systems including Linux, MacOS, Windows.
User terms: TPD is publicly accessible and non-commercial, and only provided for academic use. TPD does not reserve any uploaded data. It cleans the data and analysis results immediately the user closes the webpage. There is no user data retention time.
Current release: Release 5.0, Jan. 17, 2021
(a) Add 'Cell Fate Commitment' section.
(b) Real-time crawling the latest data and visualizing the results of TPD in COVID-19 Outbreak prediction.
(c) Add new demos and support one click to analysis and data download.
Tipping-Point Detection (TPD) is a webserver for data analysis and visualization of tipping point detection. More about TPD

TPD: Tipping-Point Detection.
The time evolution or dynamic change of many biological systems, such as disease progression and cell differentiation process, is not always smooth but occasionally abrupt, that is, there is a tipping-point or critical state during such a process at which the system state shifts irreversibly from one state (e.g. normal state) to another state (e.g. disease state), just before the critical transition. It is challenging and also important to predict such a critical state with the measured omics data, which is a key to achieve the predictive or preventive medicine. In this summary, we introduce a web service, the Tipping-Point Detection (TPD), developed to effectively and rapidly identify the tipping point during the dynamical process of biological systems, and further its leading molecules or network. The web service is based on our computational methods, landscape network entropy (LNE), sample-based local network entropy (SNE), and single-cell graph entropy, which are all model-free data-driven approaches with solid theoretical background, i.e., the dynamic network biomarker (DNB). Specifically, TPD is user-friendly and capable to explore the criticality of the dynamics from high-dimensional omics-data in terms of network entropy, thereby capturing not only the early-warning signals of the impending transition but also its leading network (DNB), which are intuitively a group of strong correlated and fluctuated molecules.
The details of the web service are as follows: The input of TPD should be the time-series or stage-course data (bulk expression matrix or single-cell data matrix) in CSV, XLS, TXT files. In the web tool, there is a flexible option for users to assign samples to certain time points. The web tool outputs multifarious visualized results including the quantitative network entropy indices (e.g., SNE scores) and their curves with suggested critical points, the identified key genes (or leading genes) and their information, DNB (the leading network that may drive the critical transition), the dynamical process of the leading network, the survival analysis based on the network entropy score that may help to identify “dark” genes (non-differential in terms of expression but differential in terms of network entropy, which may play important roles during the dynamic evolution). Thus, the web tool offers significantly enhanced display of the data and results.
Browser compatibility: TPD web tool is user-friendly for common browsers in operation systems including Linux, MacOS, Windows.
User terms: TPD is publicly accessible and non-commercial, and only provided for academic use. TPD does not reserve any uploaded data. It cleans the data and analysis results immediately the user closes the webpage. There is no user data retention time.
Citation
Last updated on Jan. 20, 2021Pei Chen, Jiayuan Zhong, Kun Yang, Xuhang Zhang, Yingqi Chen, Rui Liu* . TPD: a web tool for tipping-point detection based on dynamic network biomarker . Briefings in Bioinformatics, 2022, 23(5): bbac399.
Theoretical Background and Methods
Last updated on Jan. 20, 2021Click Methods to view.